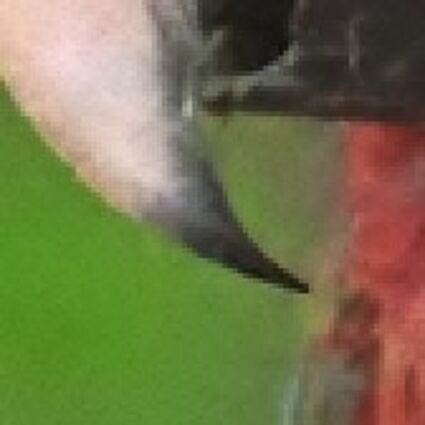
Demosaicking of noisy bayer data
Algorithm for derivation of color images from noisy Bayer-CFA data.
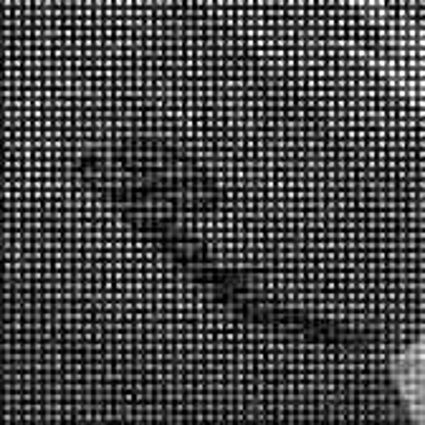
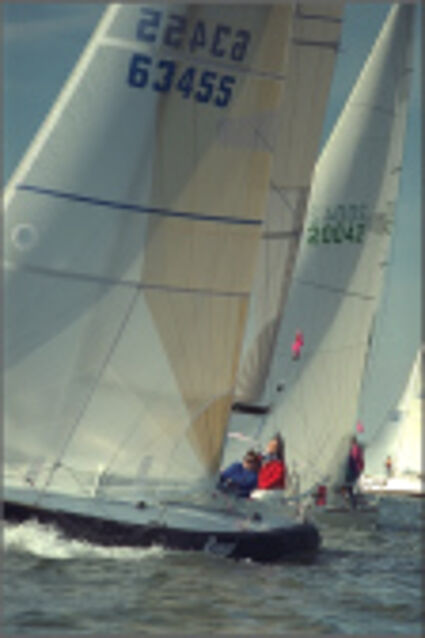
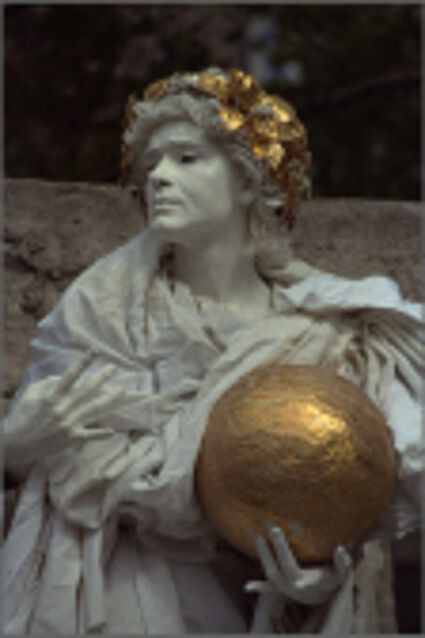
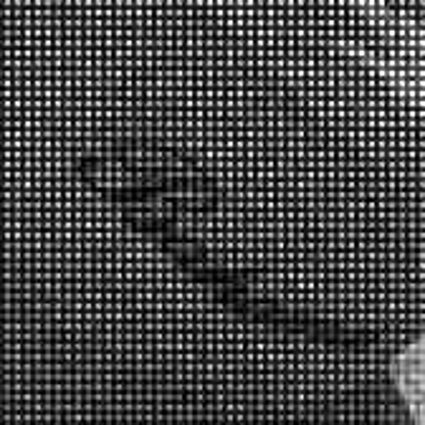
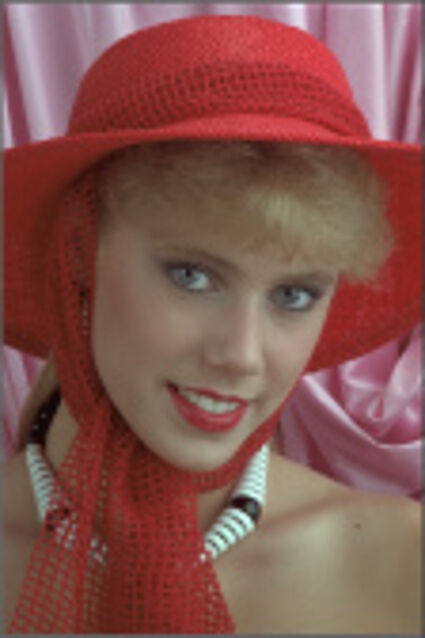
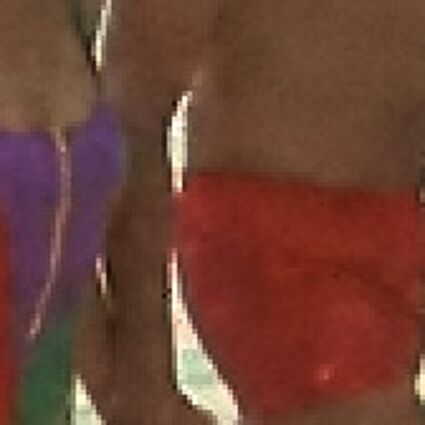
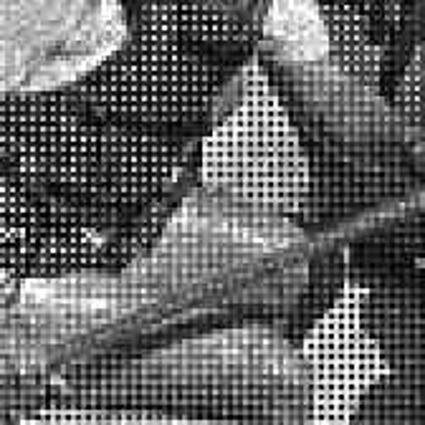
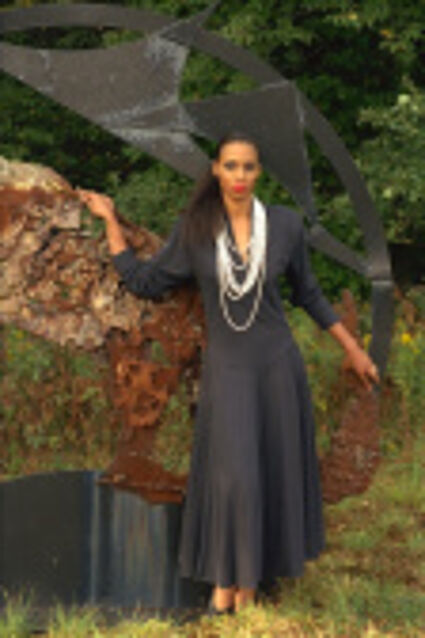
Color images usually are obtained be combining three color planes, a red, a green and a blue one. Most consumer cameras today derive color images in that way. But these cameras have only one CMOS- or CCD-sensor, to reduce the costs of production and so make the cameras cheaper. But how can one derive color images from a camera with only one sensor? The usual way is to use a color filter array (CFA), especially the Bayer color filter array as in [1]. This filter connects one of the three colors red, green or blue with the pixel coordinates of the sensor. That means that only one of three color components can pass the filter and reach the sensor plane at each pixel position. And so the problem comes up to restore the two missing color components in order to derive the expected color image. This restoring procedure is referred as demosaicking due to the mosaic like structure of the images received from the sensor.
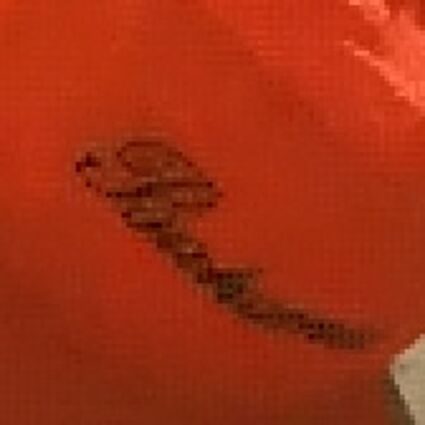
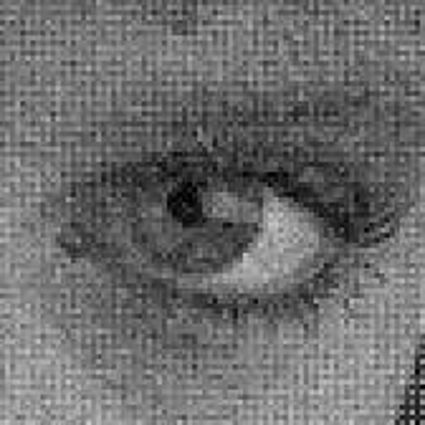
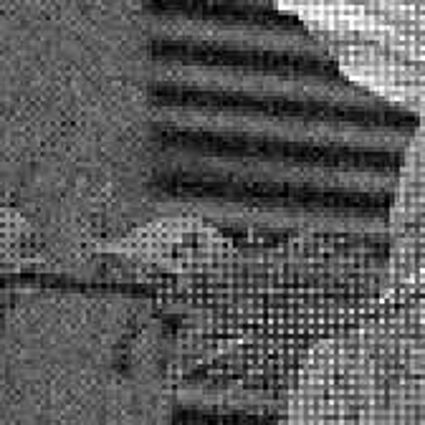
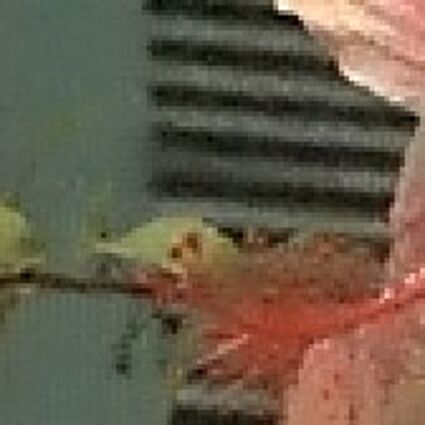
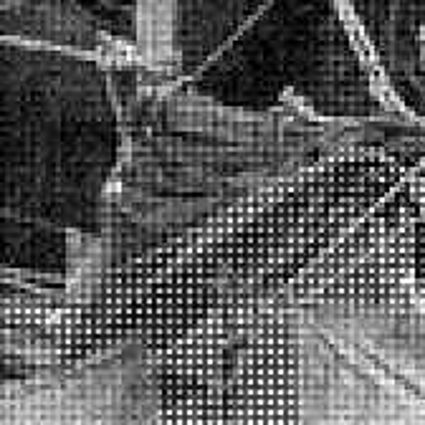
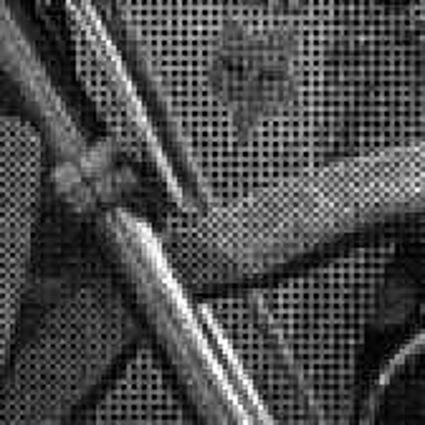
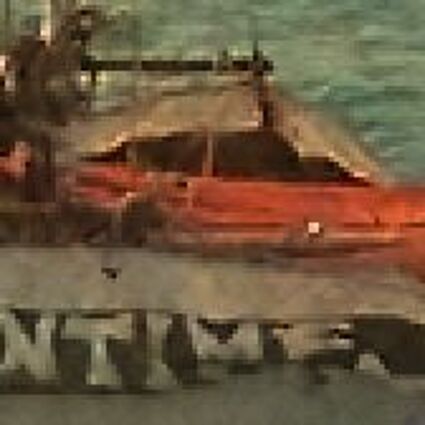
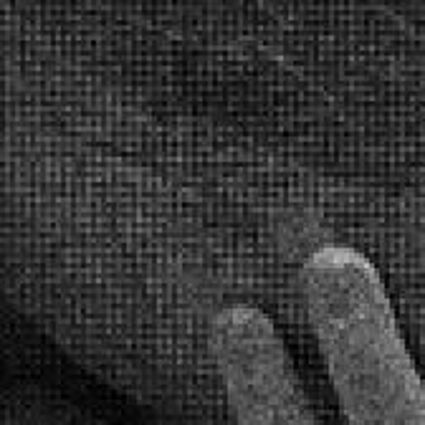
When restoring the color information from the Bayer-CFA data some typical artifacts can occure, especially on high frequency regions of images. These are known as zippering (aliasing), misguidance color artifacts and interpolation artifacts as defined in [5]. Modern algorithms can restore color information from the Bayer-CFA data while suppressing such artifacts effectively. But when noise contaminates the data most algorithms fail and these common artifacts can be seen in the resulting images.
The objective of this project is to develop algorithms which are able to restore the color information from noise free and the noisy bayer CFA data. Different noise models (e.g. Gaussian, Poisson noise) are considered. The main idea is to use non-local means filter in the noise-reduction steps.
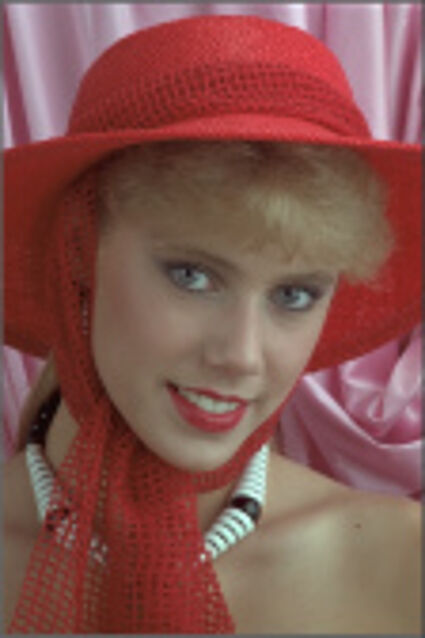
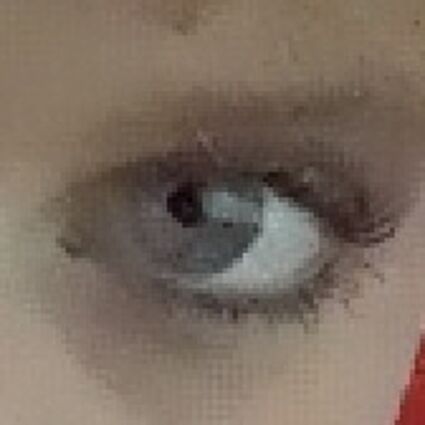
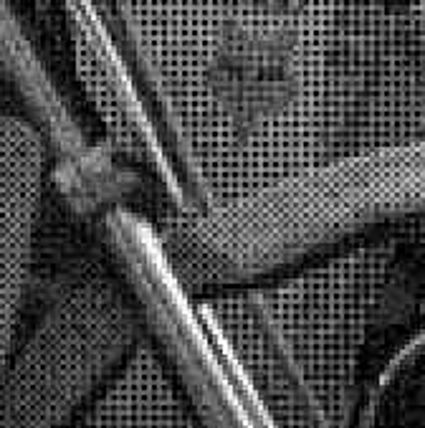
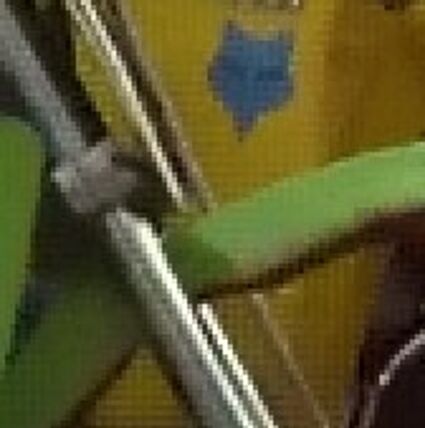
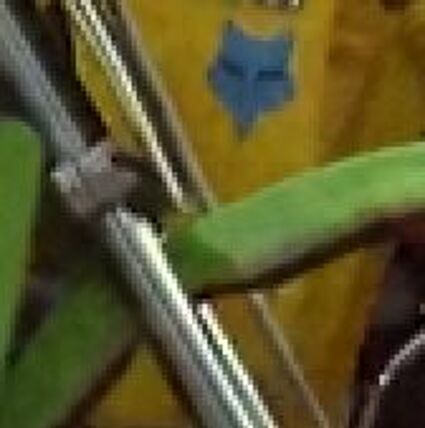
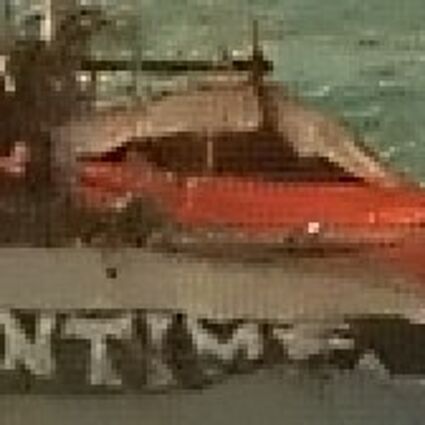
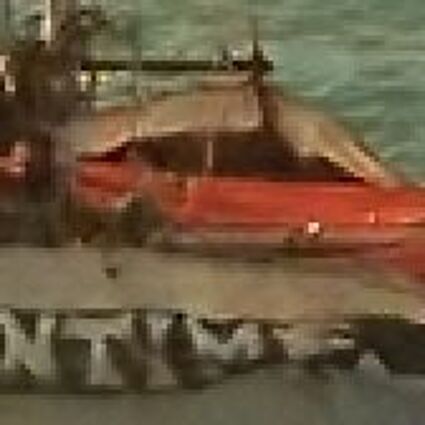
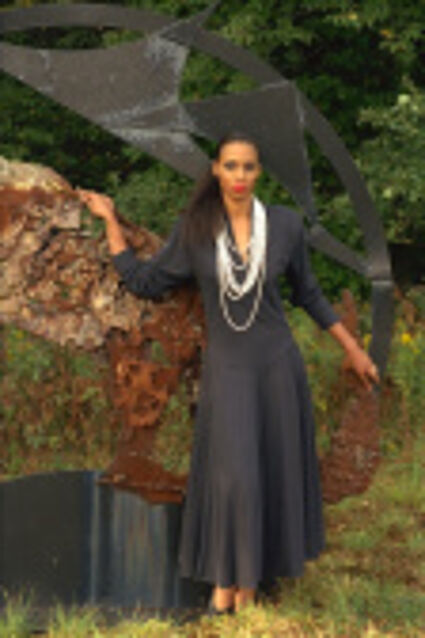
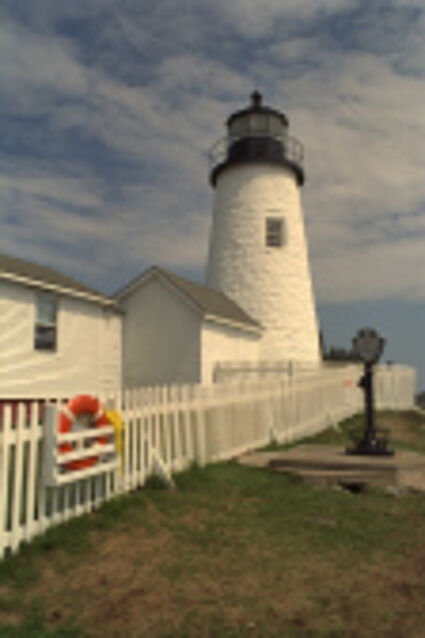
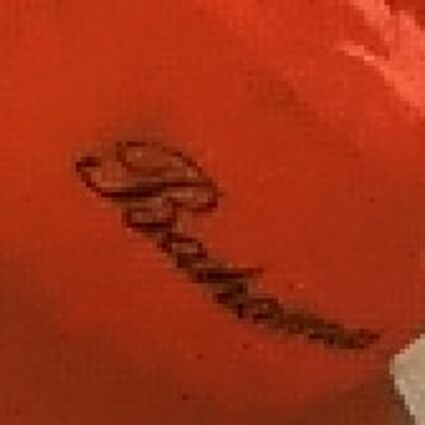
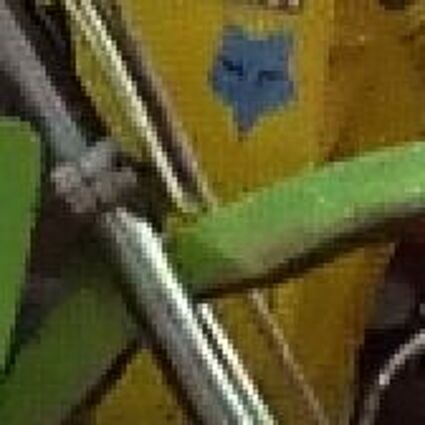
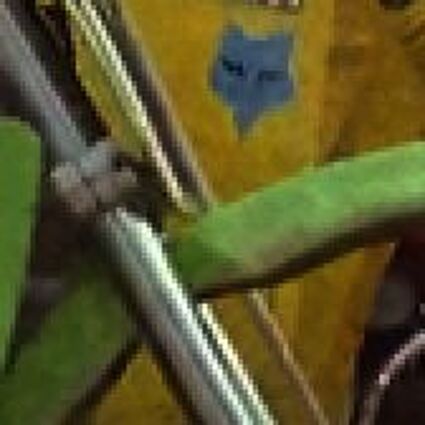
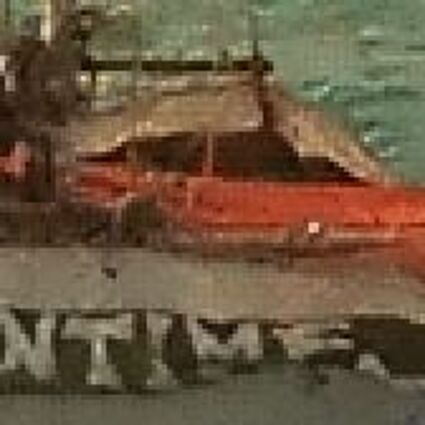
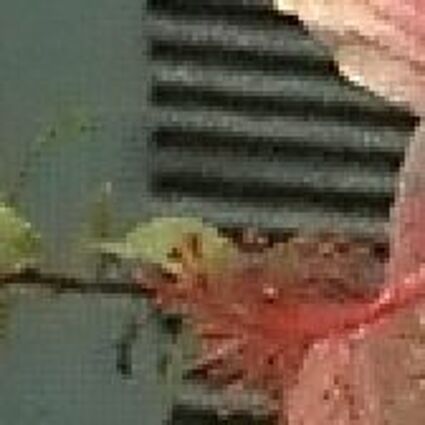
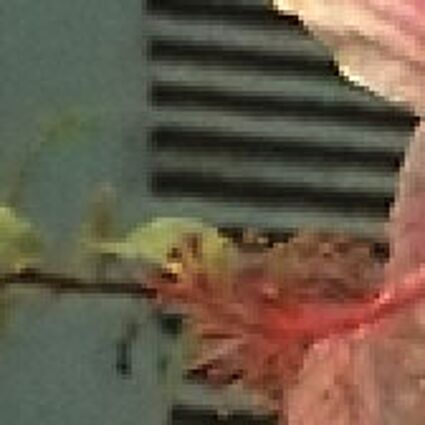
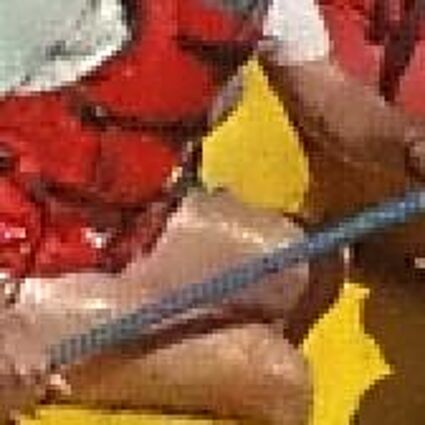
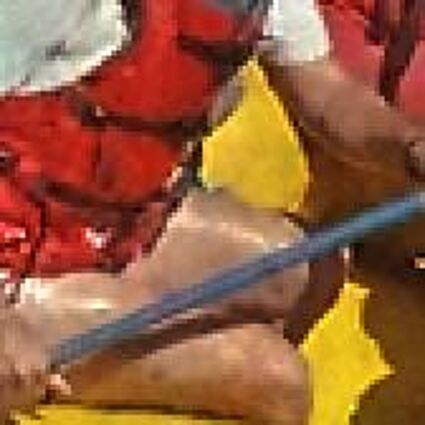
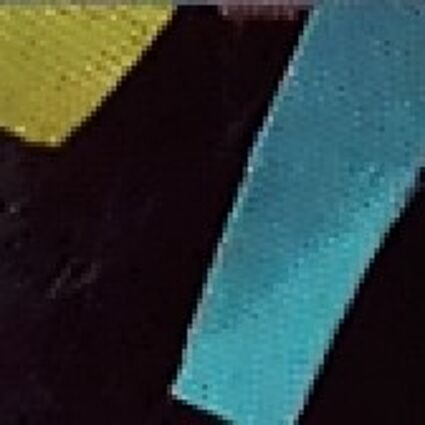
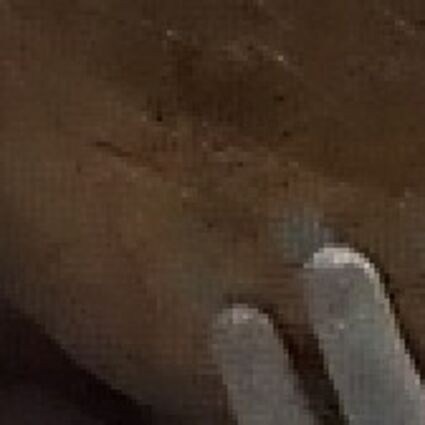
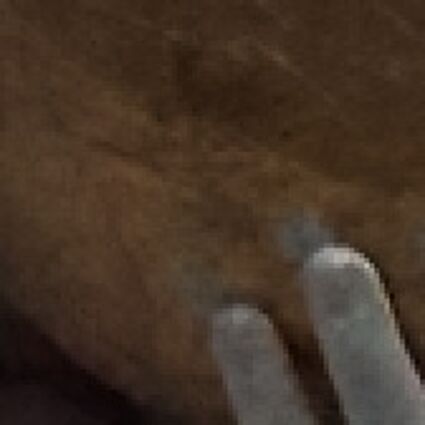
The following selected results for Gaussian and Poisson noise obtained by our algorithm described in [10] are compared with results of the algorithm in [8].
- [1]
B. E. Bayer. Color imaging array. U.S. Patent 3,971,065, 1976.
- [2]
A. Buades, B. Coll, and J.-M. Morel. Image denoising by non-local averaging. Proc. 2005 IEEE Computer Society Conference on Computer Vision and Pattern Recognition (CVPR’05), 2:60 – 65, 2005.
- [3]
A. Buades, B. Coll, J.-M. Morel, and C. Sbert. Self-similarity driven color demosaicking. IEEE Transactions on Image Processing, 18(6):1192 – 1202, 2009.
- [4]
P. J. Burt and E. H. Adelson. The laplacian pyramid as a compact image code. IEEE Transactions on Communications, 31:532–540, 1983.
- [5]
K. Hirakawa and T. W. Parks. Adaptive homogeneity-directed demosaicing algorithm. IEEE Transactions on Image Processing, 14(3):360–369, 2005.
- [6]
K. Hirakawa and T. W. Parks. Joint demosaicing and denoising. IEEE Transactions on Image Processing, 15:2146–2157, 2006.
- [7]
Y.-L. Liu, J. Wang, X. Chen, Y.-W. Guo, and Q.-S. Peng. A robust and fast non-local means algorithm for image denoising. J. Computer Science and Technology, 23(2):270–279, 2008.
- [8]
D. Paliy, V. Katkovnik, R. Bilcu, S. Alenius, and K. Egiazarian. Spatially adaptive color filter array interpolation for noiseless and noisy data. Int. J. on Imaging System Technology, 17:105–122, 2007.
- [9]
L. Zhang and X. Wu. Color demosaicking via directional linear minimum mean square-error estimation. IEEE Transactions on Image Processing, 14(12):2167–2178, 2005.
- [10]
M. Nawrath and J. Jäkel. Deriving Color Images from noisy Bayer Data using local Demosaicking and non-local Denoising 4th International Congress on Image and Signal Processing (CISP’11), Shanghai, China, 15.-17.10.2011
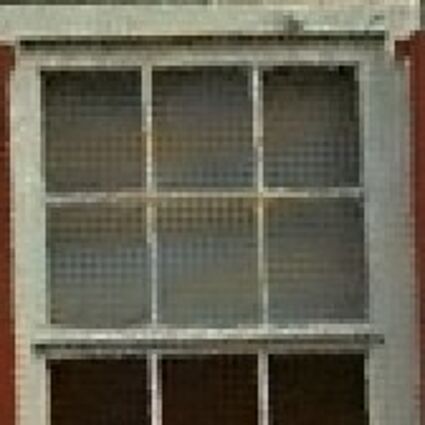
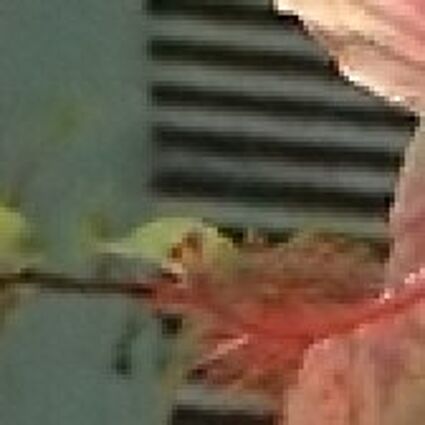
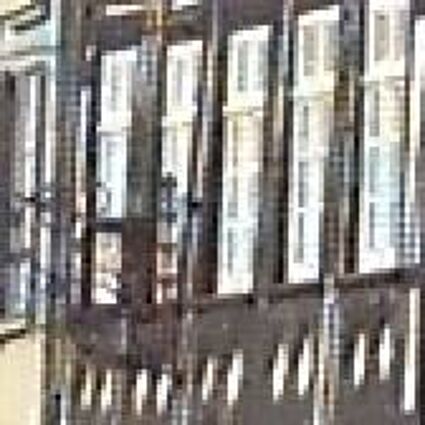
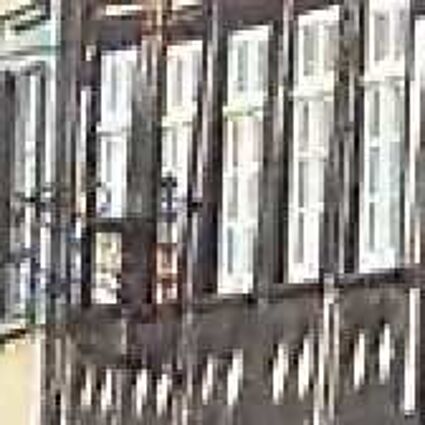
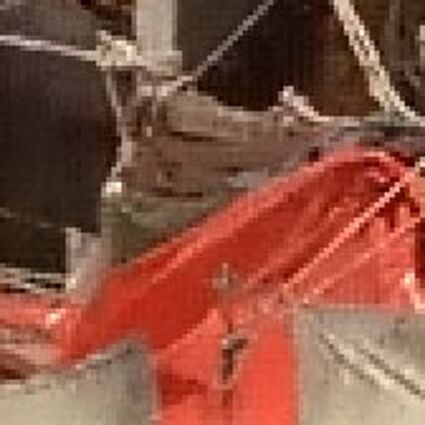
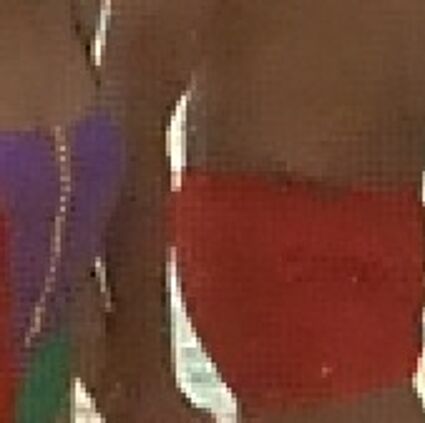
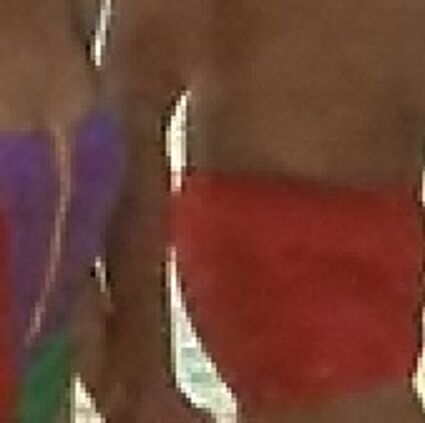
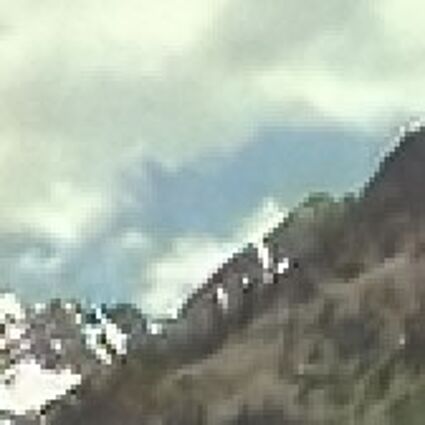
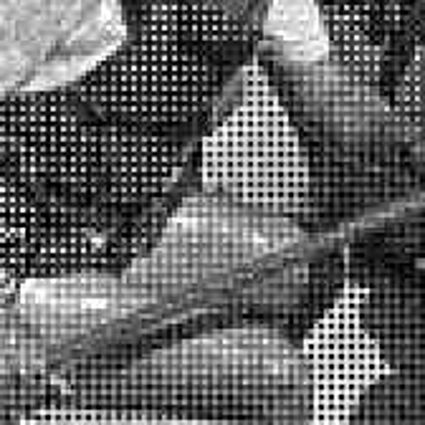
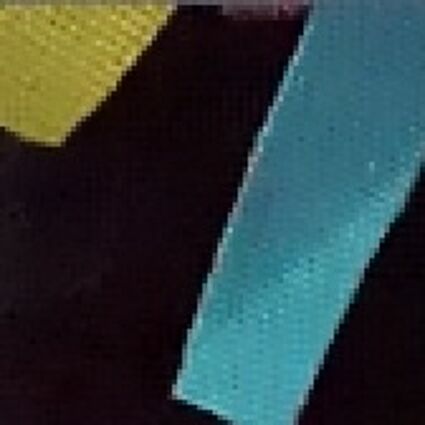
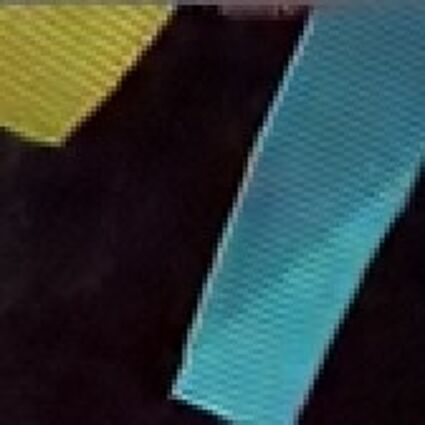
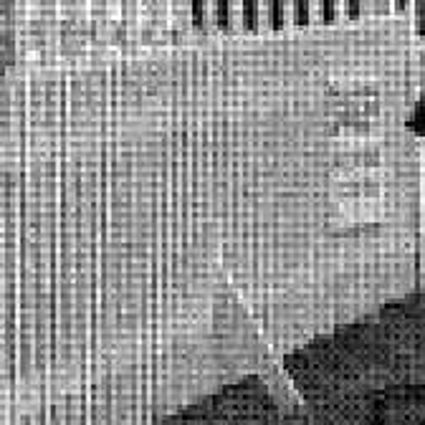
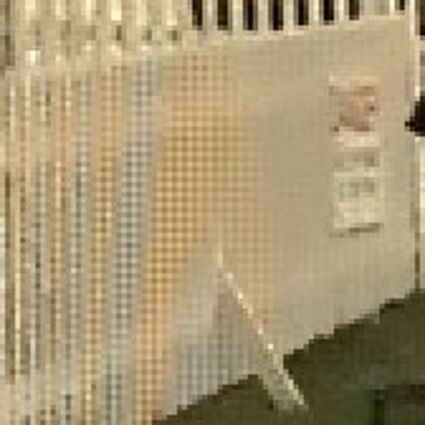
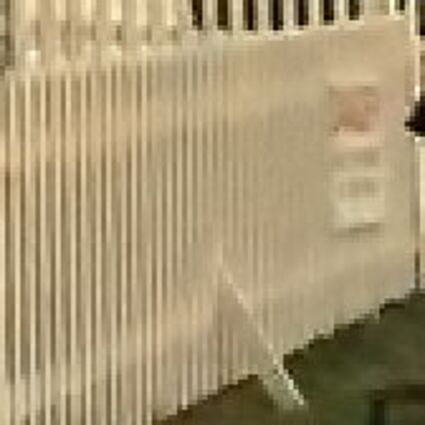
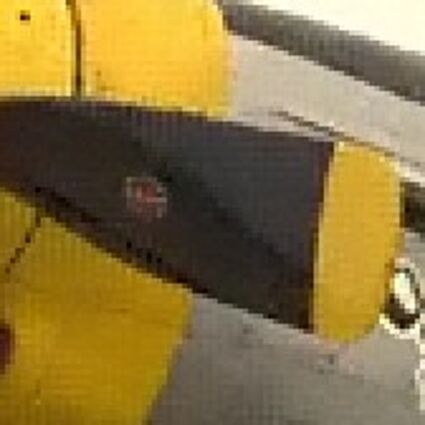
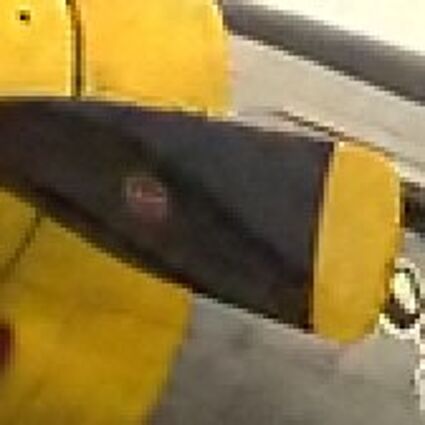
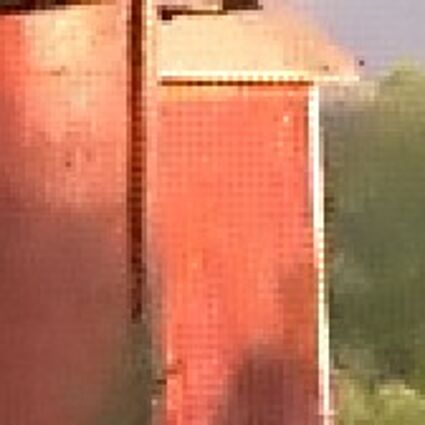
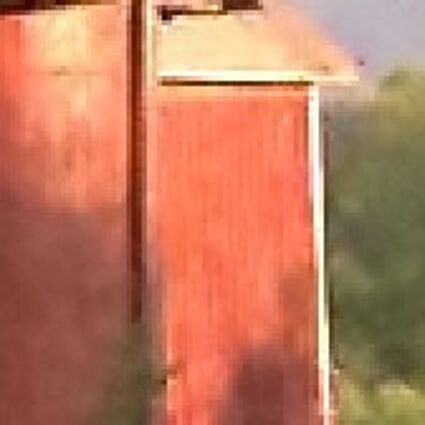
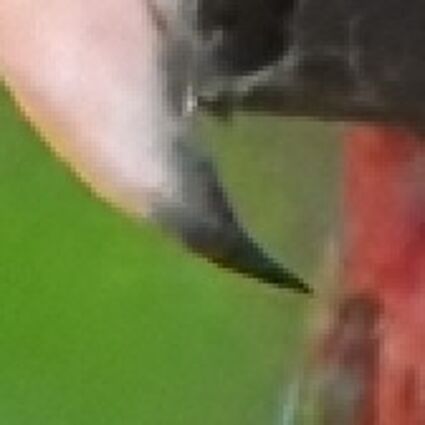
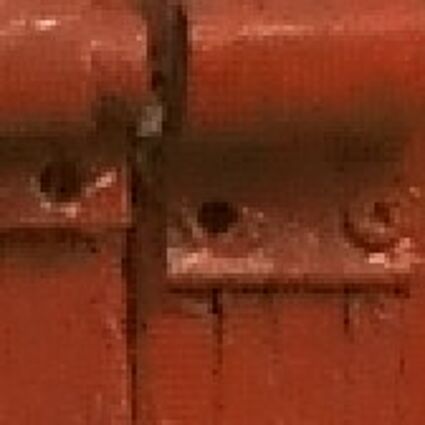
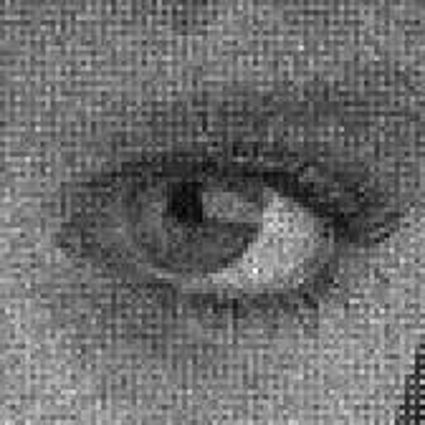
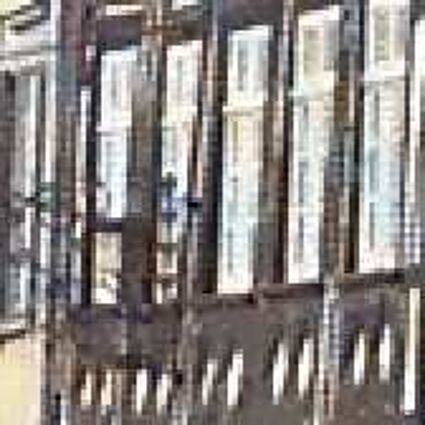
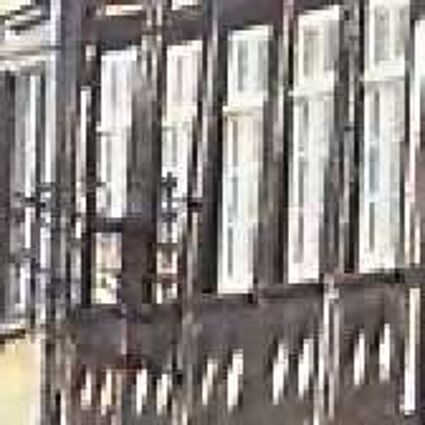
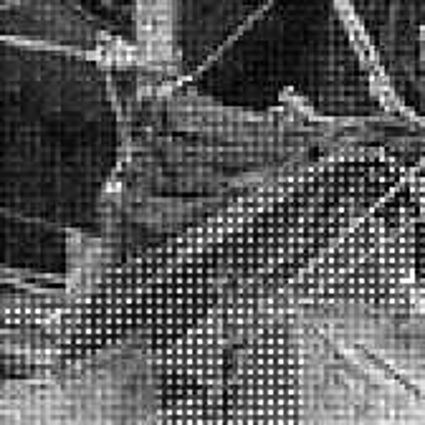
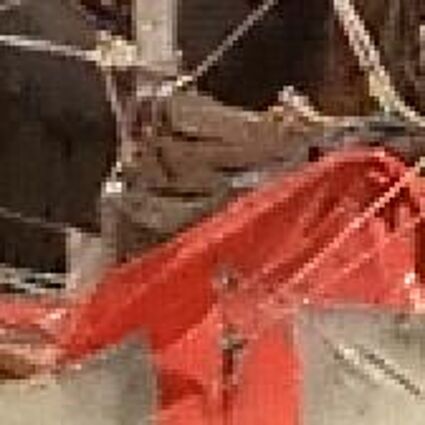
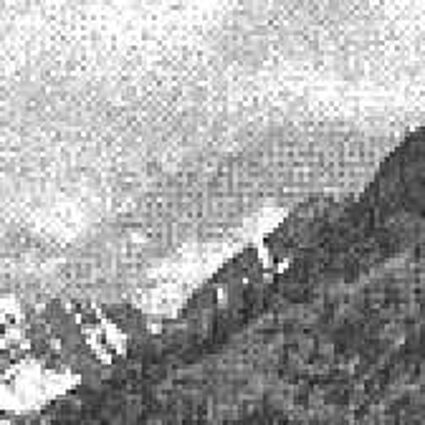
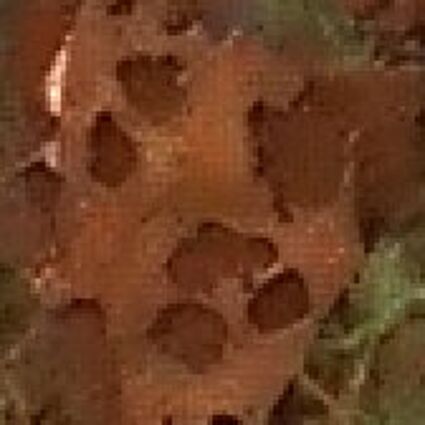
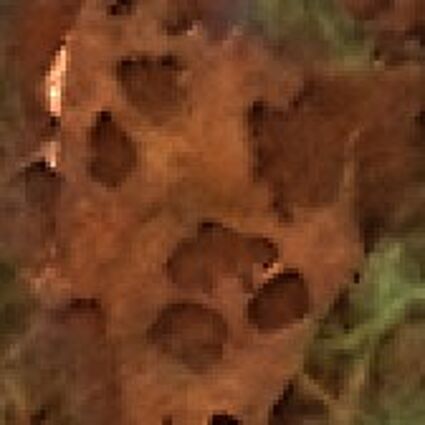
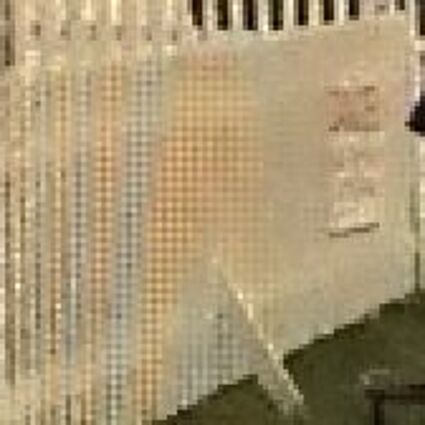
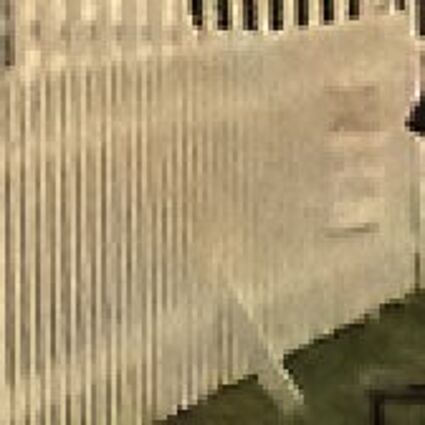
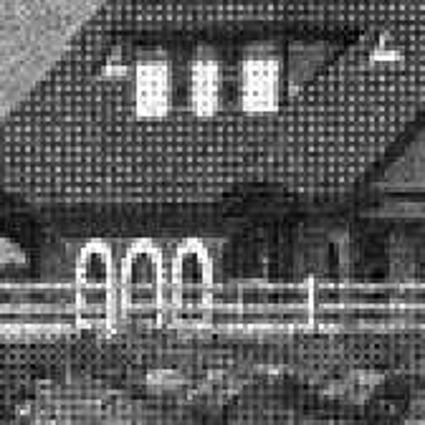
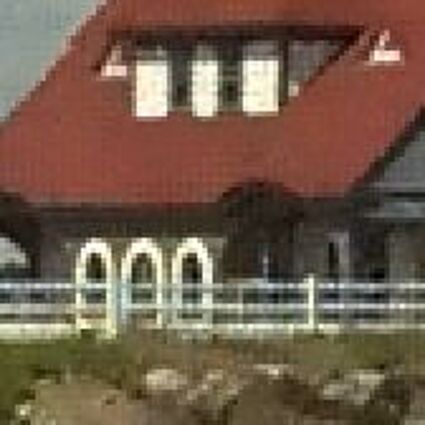
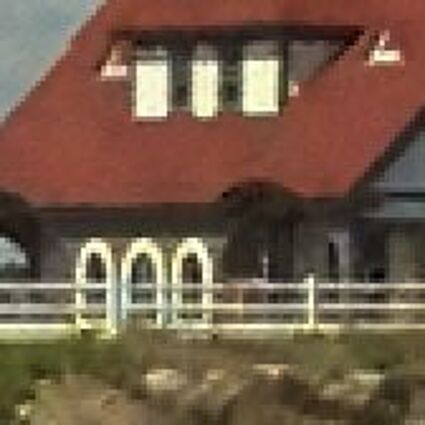
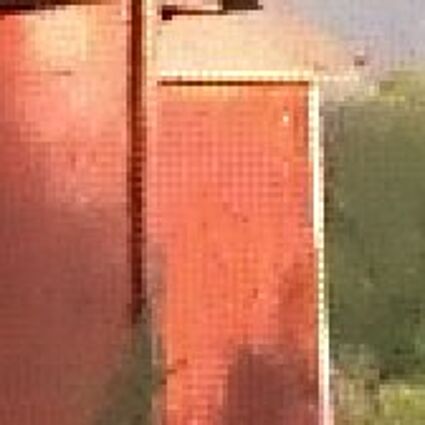
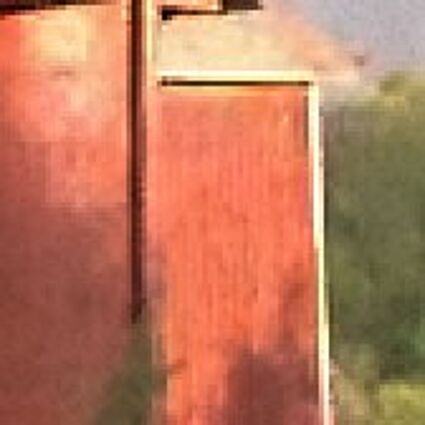
Results for gaussian noise contaminated Bayer-CFA data
The following table contains the results of the algorithm in [8] and the proposed one using Bayer-CFA data which has been contaminated with gaussian noise with a standard deviation of σ=12.75. One can click on the images to view the complete images. The images behind the links are stored as png-files to avoid jpeg-artifacts. So the sizes of the images are from about 300kB up to approximately 800kB.




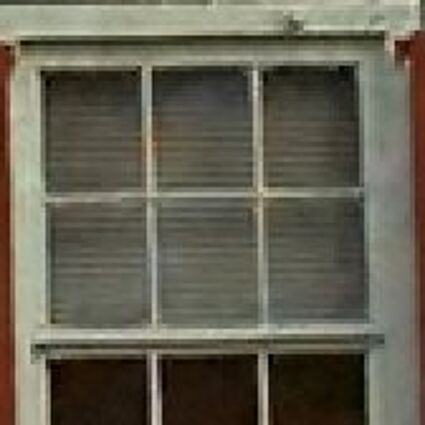
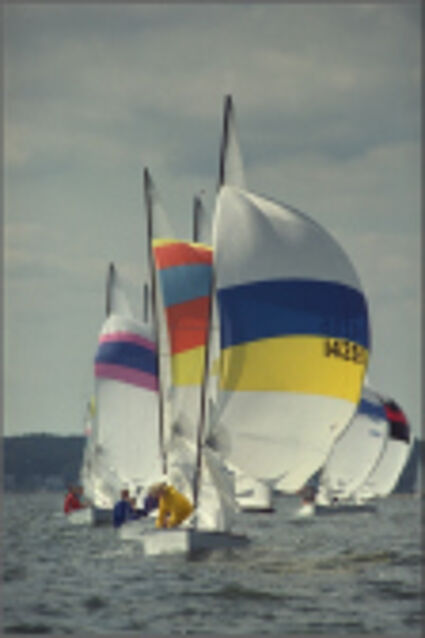
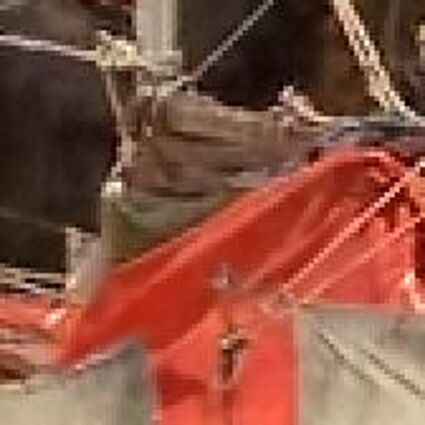
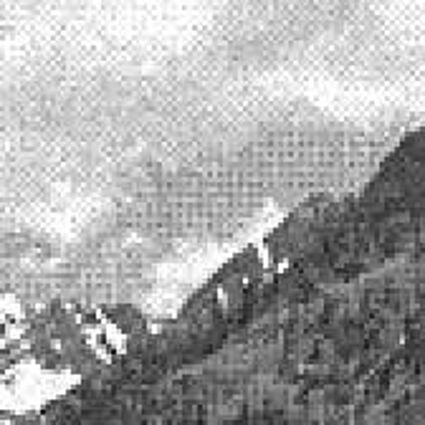
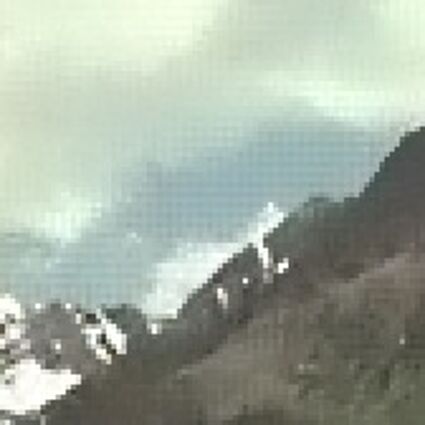
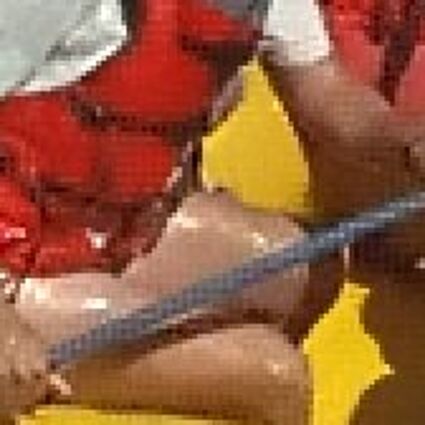
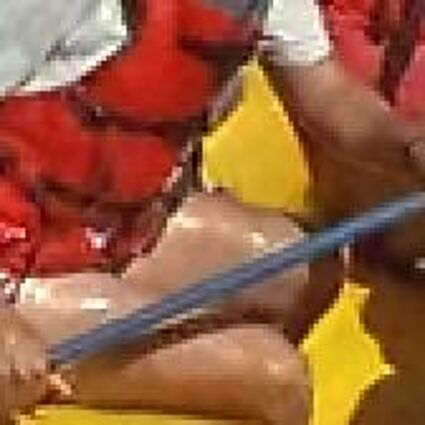
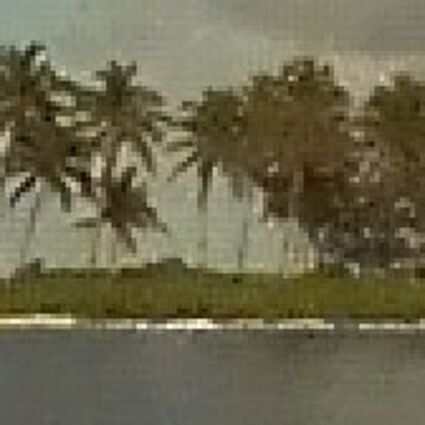
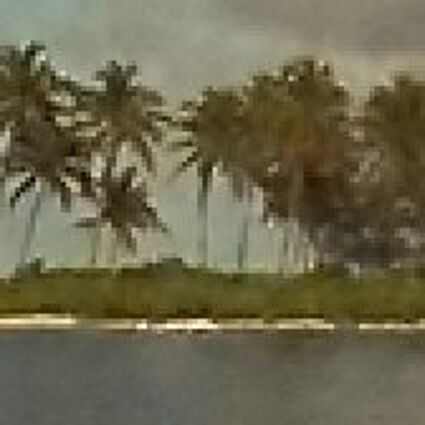
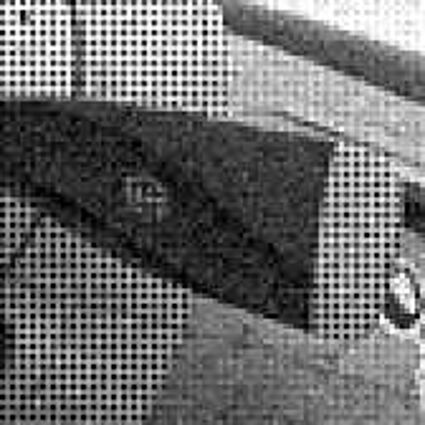
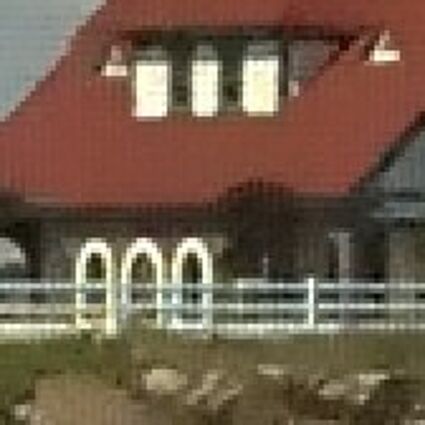
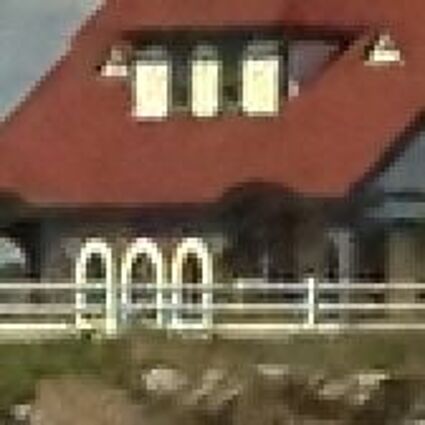
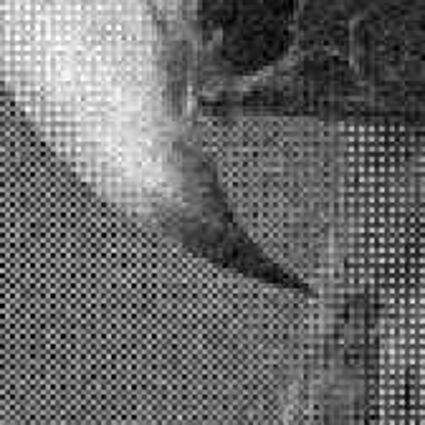
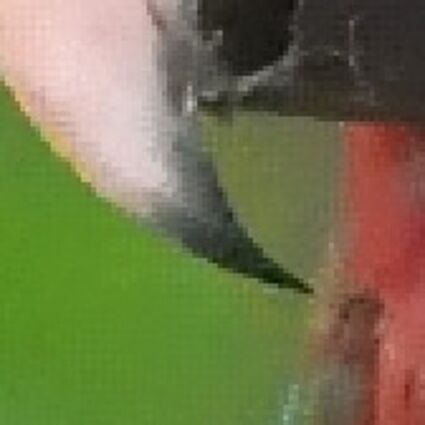
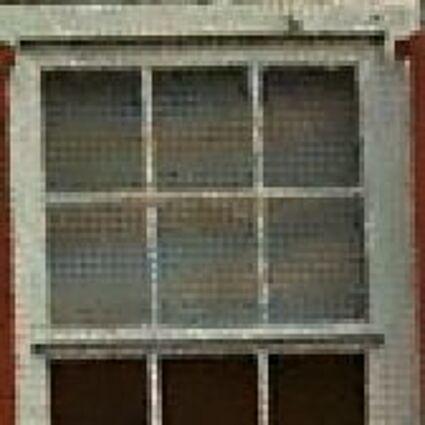
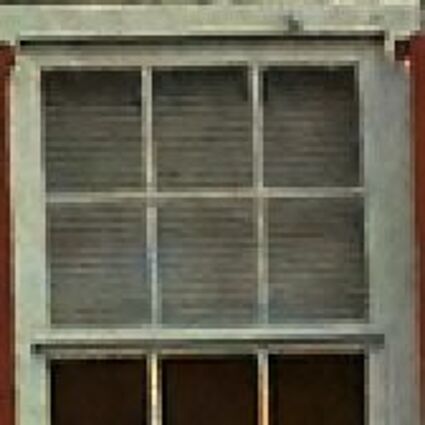
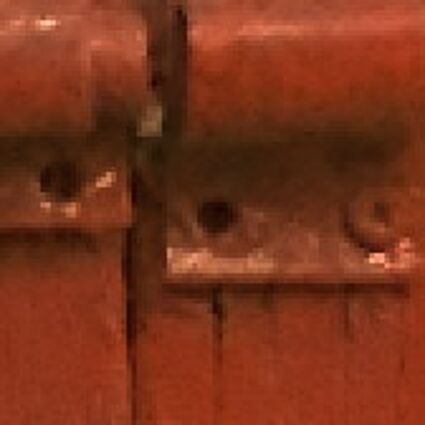
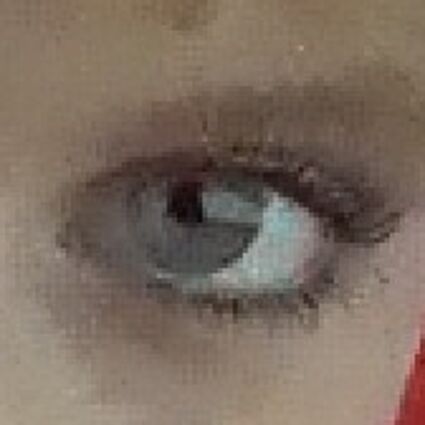
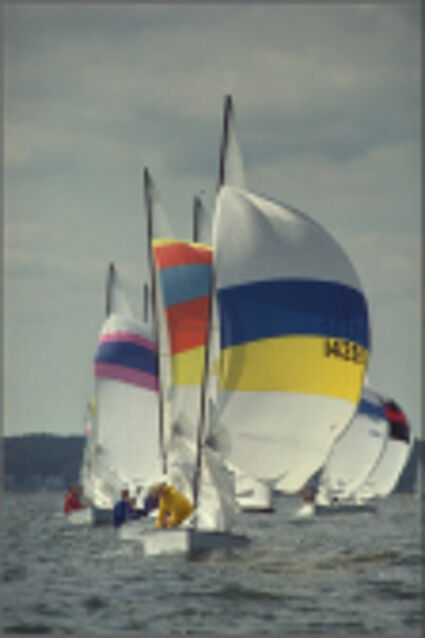
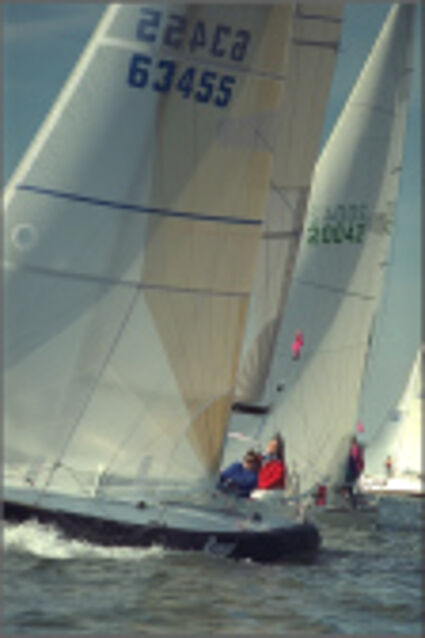
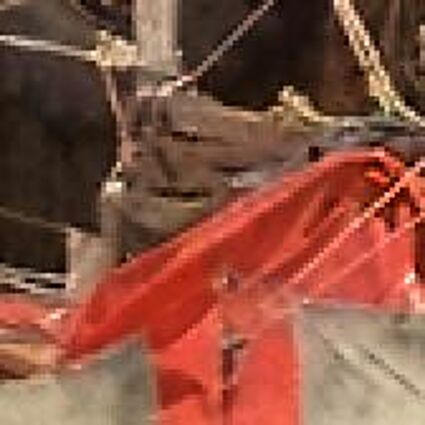
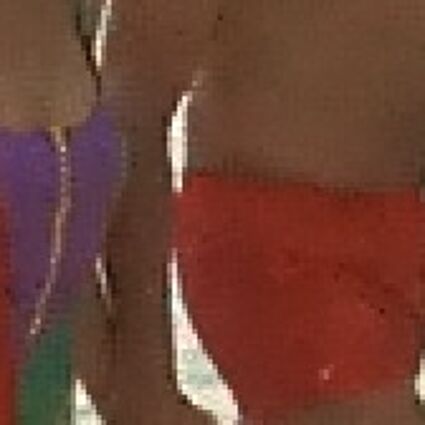
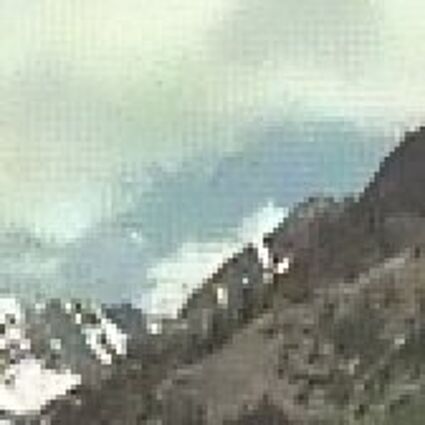
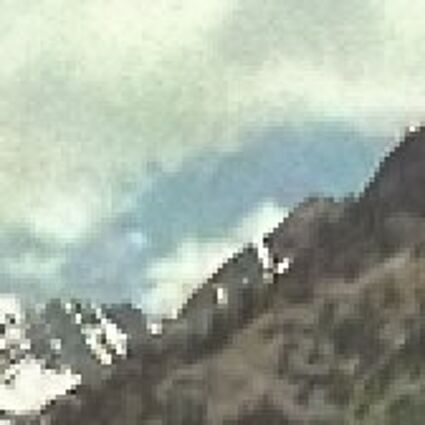
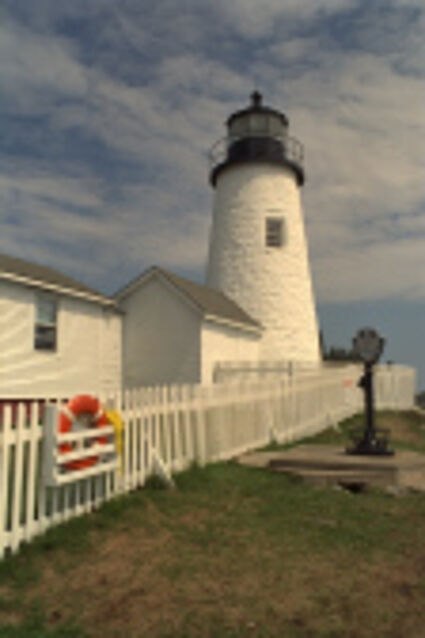
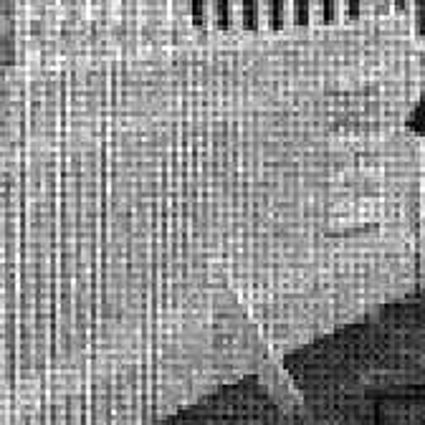
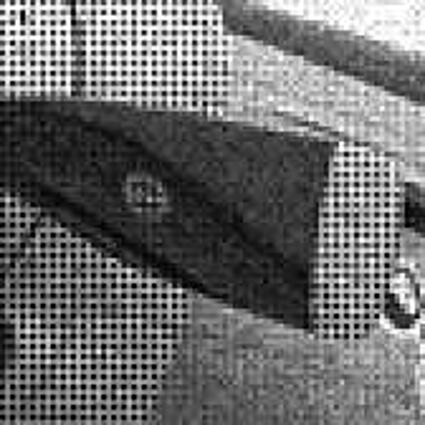
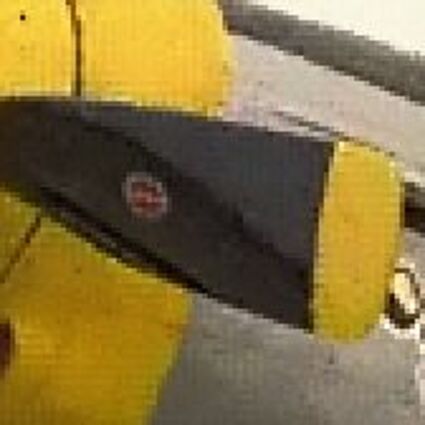
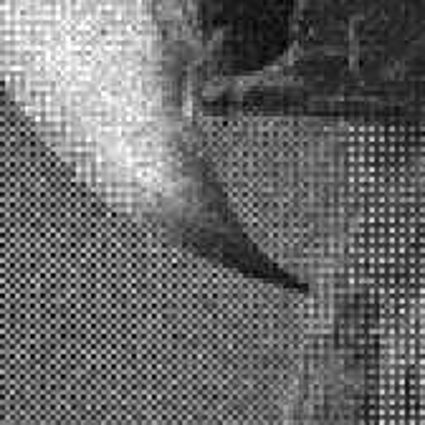
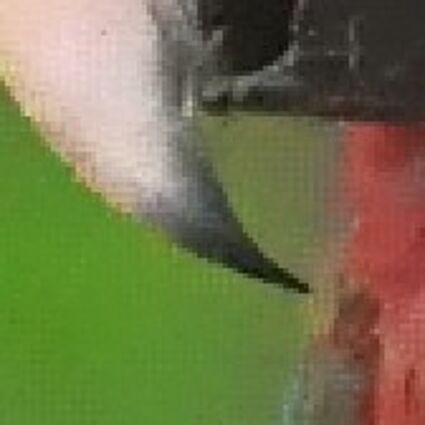
Results for Poisson noise contaminated Bayer-CFA data
The following table contains the results of the algorithm in [8] and the proposed one using Poisson noise contaminated Bayer-CFA (χ=0.5447) as source. Again, one can click on the thumbnails to view the complete images. The images behind the links are also stored as png-files to avoid jpeg-artifacts.





R
G
B